
Sorry, salad lovers. Unless you like it with a side of E. coli, romaine lettuce simply isn’t on the menu right now.
Yes, the E. coli outbreak spread by romaine lettuce that started on April 13 still isn’t fully contained, according to the Centers for Disease Control’s (CDC’s) latest update. Thanks to advances in genetic sequencing, though, we may soon have the ability to end food-borne outbreaks way faster — maybe even before anyone gets sick.
We haven’t always been great at addressing outbreaks of food-borne illness. In the past, we typically only knew something was wrong when a lot of people got sick at the same time or in the same area. Investigators would interview these people, looking for connections between them — maybe they all ate food from the same restaurant, or bought the same brand of groceries.
This approach was both time-consuming and imprecise.

In the 1990s, researchers turned to genetics to help investigate food-borne outbreaks. They’d analyze clumps of DNA from microbes gathered from the people who had gotten ill using a technique called pulsed-field gel electrophoresis. This let researchers create a “fingerprint” for various bacterias.
If samples from two or more ill people contained germs with the same genetic fingerprint, researchers would know their cases shared a connection. They could then look for the same fingerprint in food samples, from a restaurant or processing plant, for example. Find another match, and they’d know they’d found their outbreak source.
This worked better than just asking people. But it was still time-consuming and could miss related cases, according to the CDC.
Then, in 2013, investigators across the nation began using whole genome sequencing to look at a microbe’s entire DNA profile, not just bits of it.
Whole genome sequencing is more precise than other methods, and it just keeps getting faster and cheaper. Joel Sevinsky, head of the Molecular Science Laboratory at the Colorado Department of Public Health and Environment (CDPHE), told the Associated Press his lab can sequence the genome of a suspected pathogen in less than 72 hours.
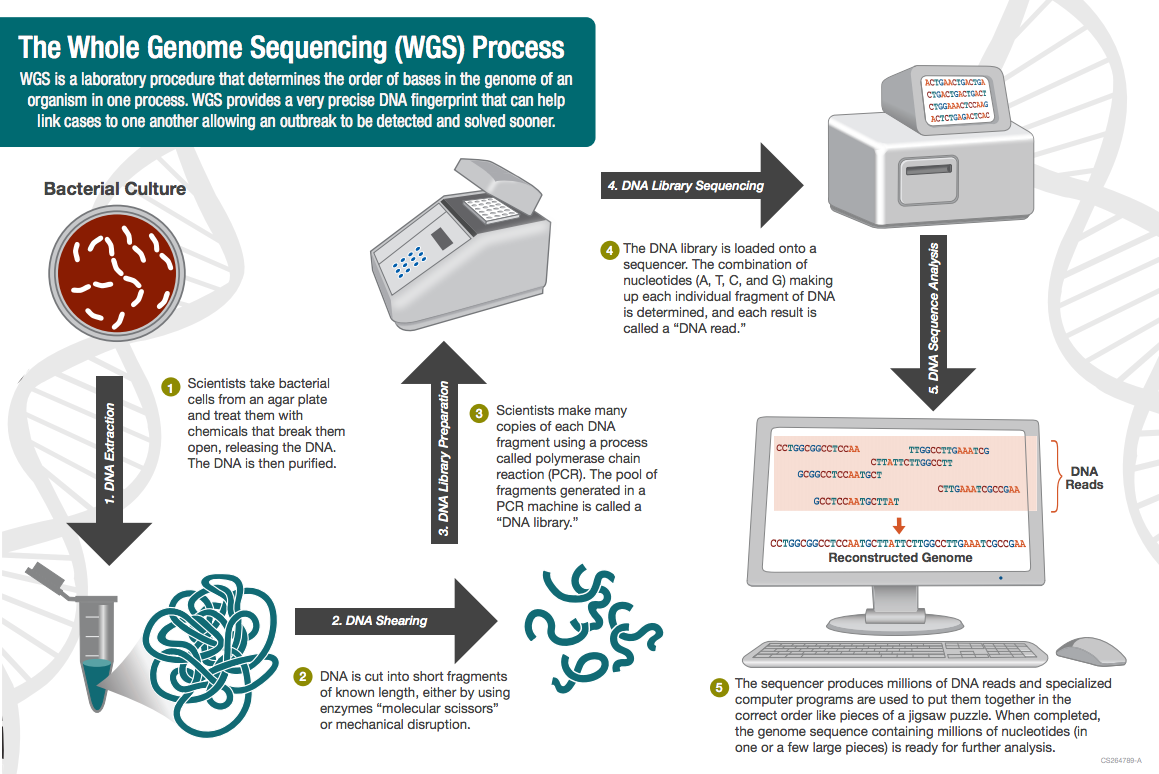
Whole genome sequencing is already helping researchers address food-borne outbreaks, including a 2017 salmonella outbreak that stretched across 21 states, and the current romaine outbreak.
It’s even identifying contaminated food before it even reaches the public. According to the AP, inspectors used genome sequencing to find pathogens that could have caused outbreaks when they inspected food plants. They were able to recall the tainted products before they ever reached grocery stores or restaurants, preventing countless people from being sickened.
Today, states without the necessary lab capabilities send their samples to the CDC lab in Atlanta for whole genome sequencing. But soon, they won’t have to.
The CDC is spending $12 million to help state and city health departments set up their own whole genome sequencing labs. By the end of 2018, they expect every state will be able to conduct its own tests, so they can track pathogens from salmonella to E. coli.
We couldn’t avoid this romaine outbreak, it seems. But with more tools for detection, we might be able to prevent the next one.